PAM50 Template¶
Introduction¶
Template-based analysis of multi-parametric MRI data of the spinal cord sets the foundation for standardization and reproducibility. Particularly, it allows researchers to process their data with minimum bias and perform multi-center studies, with the goal of improving patient diagnosis and prognosis and discovering new biomarkers of spinal-related diseases.
To perform template-based analysis, a template is needed. Templates act as a common reference space to realign each subject to. PAM50 is one such template for MRI of the full spinal cord and brainstem, and is included in your installation of SCT. “PAM” stands for “Polytechnique”, “Aix-Marseille University” and “Montreal Neurological Institute”, the three institutions that collaborated to develop the template. The PAM50 template has the following features:
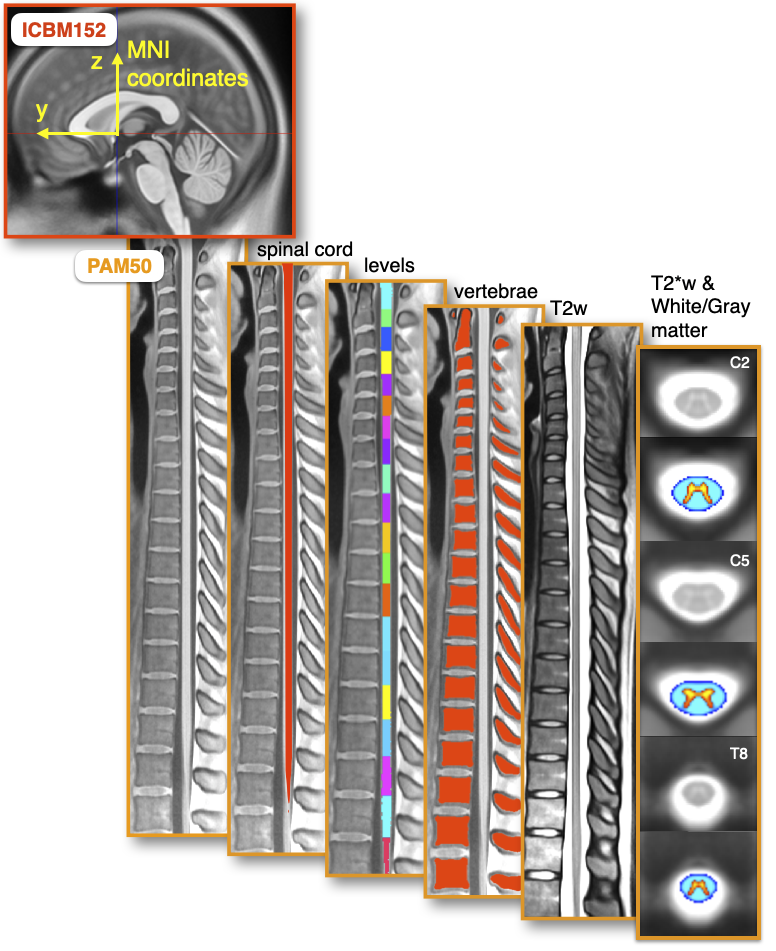
It is available for T1-, T2-and T2*-weighted MRI contrast.
It is compatible with the ICBM152 brain template (a.k.a. the MNI coordinate system), allowing researchers to conduct simultaneous brain/spine studies within the same coordinate system.
It includes a probabilistic atlas for white matter, gray matter, and CSF regions.
It is a straightened template rather than a curved one.
The template is stored in the data/PAM50/ directory in your SCT installation directory.
PAM50/ contains several subfolders, each corresponding to different aspects of PAM50.
Note
For more details on the creation of the template, see the neuropoly/template GitHub repository.
Template¶
Location: PAM50/template
The template folder contains the PAM50 template for various MRI contrast types. It also contains binary and probabilistic masks for different spinal features, as well as point-wise labels for vertebral levels and intervertebral discs.
The template folder also contains an info_label.txt file to explain what each file represents:
# Template labels
# Keyword=IndivLabels (Please DO NOT change this line)
# ID, name, file
0, T1-weighted template, PAM50_t1.nii.gz
1, T2-weighted template, PAM50_t2.nii.gz
2, T2S-weighted template, PAM50_t2s.nii.gz
3, spinal cord mask (binary), PAM50_cord.nii.gz
4, white matter mask (probabilistic), PAM50_wm.nii.gz
5, gray matter mask (probabilistic), PAM50_gm.nii.gz
6, cerebrospinal fluid mask (binary), PAM50_csf.nii.gz
7, spinal cord mask with discrete vertebral levels, PAM50_levels.nii.gz
8, spinal cord mask with continuous vertebral levels, PAM50_levels_continuous.nii.gz
9, point-wise vertebral body labels, PAM50_label_body.nii.gz
10, point-wise intervertebral disc labels, PAM50_label_disc.nii.gz
11, point-wise posterior intervertebral disc labels, PAM50_label_discPosterior.nii.gz
12, spine mask (binary), PAM50_spine.nii.gz
13, spinal cord centerline, PAM50_centerline.nii.gz
14, point-wise spinal level labels, PAM50_spinal_midpoint.nii.gz
15, spinal cord mask with discrete spinal levels, PAM50_spinal_levels.nii.gz
Please note that the white and gray matter masks found in template/ are for the entire structures.
If you are looking for individual tracts, the relevant volumes are provided within the White and Gray Matter Atlas section below.
White and Gray Matter Atlas¶
Location: PAM50/atlas
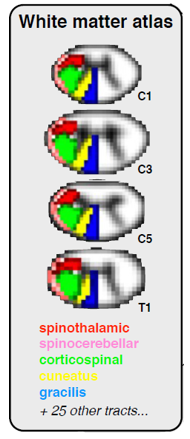
The White Matter atlas will be a useful tool for your studies of specific spinal cord tracts. It consists of 36 NIfTI volumes named PAM50_atlas_<tract_number>.nii.gz where <tract_number> is the number identifying the tract. Fifteen WM tracts and three GM regions are available for each side. The values of each voxel of the files PAM50_atlas_<tract_number>.nii.gz are the voxel volume proportions occupied by the corresponding tract.
Notably, each volume is a probabilistic mask, or “soft mask”. The voxel values in these volumes range from 0 to 1; voxels in the center of each tract will equal 1, while voxels near the edges of each tract will be weighted closer to 0. This is to account for partial volume effects.
The atlas folder also contains an info_label.txt file to explain what each file represents:
# White matter atlas. Generated on: 01-Sep-2016
#
# Keyword=IndivLabels (Please DO NOT change this line)
# ID, name, file
0, WM left fasciculus gracilis, PAM50_atlas_00.nii.gz
1, WM right fasciculus gracilis, PAM50_atlas_01.nii.gz
2, WM left fasciculus cuneatus, PAM50_atlas_02.nii.gz
3, WM right fasciculus cuneatus, PAM50_atlas_03.nii.gz
4, WM left lateral corticospinal tract, PAM50_atlas_04.nii.gz
5, WM right lateral corticospinal tract, PAM50_atlas_05.nii.gz
6, WM left ventral spinocerebellar tract, PAM50_atlas_06.nii.gz
7, WM right ventral spinocerebellar tract, PAM50_atlas_07.nii.gz
8, WM left rubrospinal tract, PAM50_atlas_08.nii.gz
9, WM right rubrospinal tract, PAM50_atlas_09.nii.gz
10, WM left lateral reticulospinal tract, PAM50_atlas_10.nii.gz
11, WM right lateral reticulospinal tract, PAM50_atlas_11.nii.gz
12, WM left spinal lemniscus (spinothalamic and spinoreticular tracts), PAM50_atlas_12.nii.gz
13, WM right spinal lemniscus (spinothalamic and spinoreticular tracts), PAM50_atlas_13.nii.gz
14, WM left spino-olivary tract, PAM50_atlas_14.nii.gz
15, WM right spino-olivary tract, PAM50_atlas_15.nii.gz
16, WM left ventrolateral reticulospinal tract, PAM50_atlas_16.nii.gz
17, WM right ventrolateral reticulospinal tract, PAM50_atlas_17.nii.gz
18, WM left lateral vestibulospinal tract, PAM50_atlas_18.nii.gz
19, WM right lateral vestibulospinal tract, PAM50_atlas_19.nii.gz
20, WM left ventral reticulospinal tract, PAM50_atlas_20.nii.gz
21, WM right ventral reticulospinal tract, PAM50_atlas_21.nii.gz
22, WM left ventral corticospinal tract, PAM50_atlas_22.nii.gz
23, WM right ventral corticospinal tract, PAM50_atlas_23.nii.gz
24, WM left tectospinal tract, PAM50_atlas_24.nii.gz
25, WM right tectospinal tract, PAM50_atlas_25.nii.gz
26, WM left medial reticulospinal tract, PAM50_atlas_26.nii.gz
27, WM right medial reticulospinal tract, PAM50_atlas_27.nii.gz
28, WM left medial longitudinal fasciculus, PAM50_atlas_28.nii.gz
29, WM right medial longitudinal fasciculus, PAM50_atlas_29.nii.gz
30, GM left ventral horn, PAM50_atlas_30.nii.gz
31, GM right ventral horn, PAM50_atlas_31.nii.gz
32, GM left intermediate zone, PAM50_atlas_32.nii.gz
33, GM right intermediate zone, PAM50_atlas_33.nii.gz
34, GM left dorsal horn, PAM50_atlas_34.nii.gz
35, GM right dorsal horn, PAM50_atlas_35.nii.gz
36, CSF contour, PAM50_atlas_36.nii.gz
# Combined labels
# Keyword=CombinedLabels (Please DO NOT change this line)
# ID, name, IDgroup
50, spinal cord, 0:35
51, white matter, 0:29
52, gray matter, 30:35
53, dorsal columns, 0:3
54, lateral funiculi, 4:13
55, ventral funiculi, 14:29
# Clusters used for the first step of the MAP estimation (for advanced users only)
# Keyword=MAPLabels (Please DO NOT change this line)
# Name, IDgroup
white matter, 0:29
gray matter, 30:35
CSF, 36
Note
For more details on the implementation of atlas, see the White matter atlas script.
Spinal levels¶
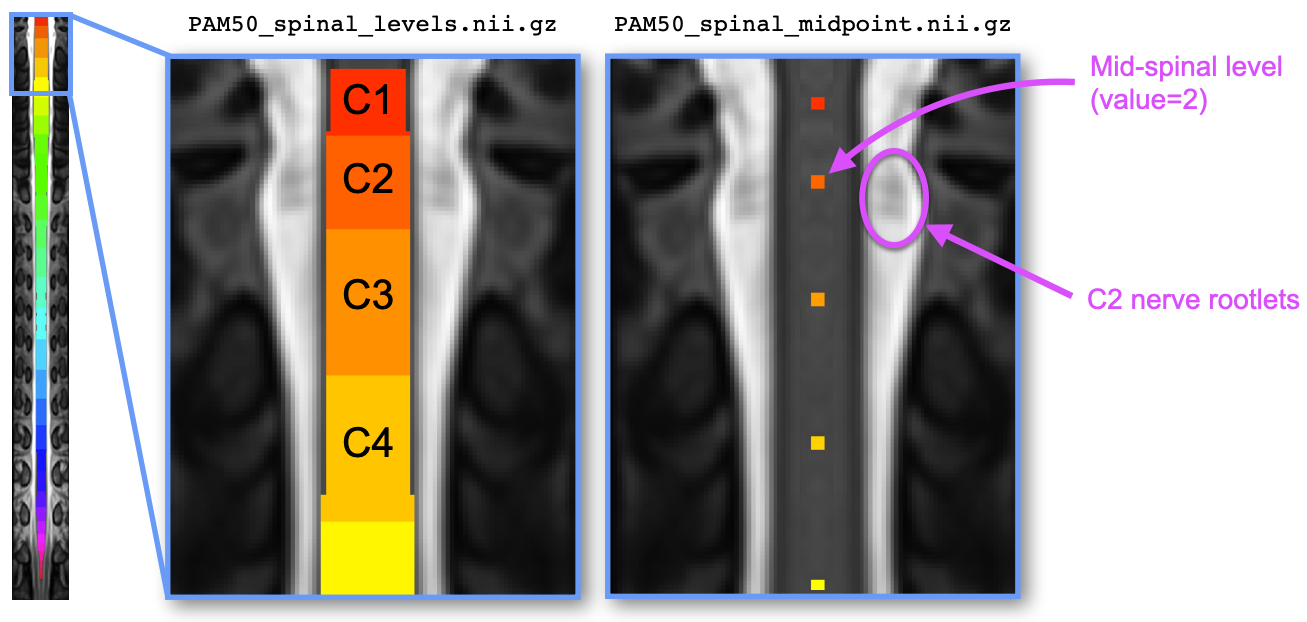
The file PAM50/template/PAM50_spinal_levels.nii.gz represents the spinal cord segmentation with parcellation
across 30 spinal levels ranging from C1 to S5. Like for the vertebral level file, each level is encoded with
an integer value (C1: 1, C2: 2, …, C8: 8, T1: 9, etc.).
The file PAM50/template/PAM50_spinal_midpoint.nii.gz represents the mid-point of the spinal level. Each level is
encoded by one voxel centered in the spinal cord, and the value of the voxel corresponds to the spinal level. This
file can be useful for registration to the PAM50 using the spinal levels instead of the intervertebral discs, if the
former are available (e.g., via nerve rootlets segmentation).
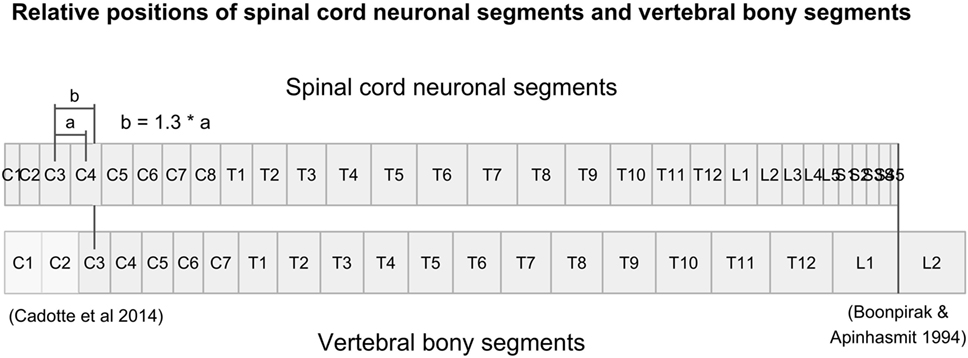
The spinal levels are estimated from the relative length of each spinal level with respect to the length of the full spinal cord (expressed as a percentage), found in Table 3 of the article by Frostell et al. (2016). The figure below (extracted from Frostell et al.) shows the spatial correspondance between the spinal vs. vertebral levels.
Note
For more details on the implementation of the spinal levels in SCT, see the Pull Request on GitHub.