Registration Option 1: Reusing previous warping fields¶
Say that you have already registered some T2 anatomical data, and you have MT data from the same session that you would also like to register to the template. In that case, there is no need to run sct_register_to_template again, because you can reuse the warping field between the template and the T2 anatomical space. This has several benefits:
It provides consistency between contrasts, as the “affine” part of the transformation (in other words, the matching between the vertebral levels of the data and template) will be the same for both contrasts.
It saves processing time, because you can skip the vertebral labeling step and part of the registration process.
For the purpose of this tutorial, we will register the MT data alongside the T2 results from the previous Registering labeled anatomical images to the PAM50 template tutorial. Specifically, we will reuse the /t2/warp_template2anat.nii.gz warping field to initialize the registration for the MT data.
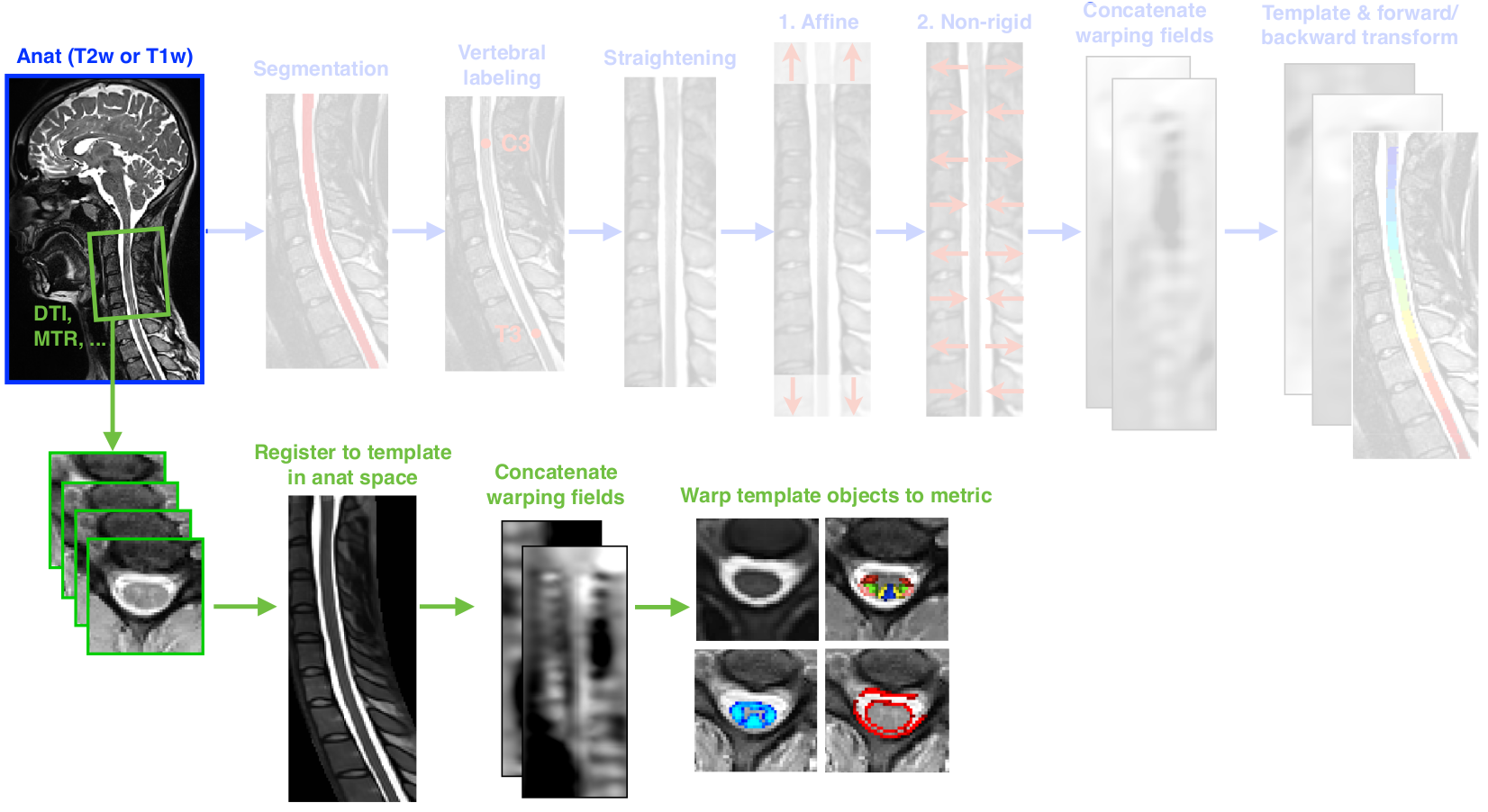
Template->T2 (blue) + T2->MT1 (green) = Template->MT1 (the desired result)¶
Since we are starting the MT registration with an initial transformation already applied, all that remains is fine-tuning for the MT data. So, here we use a different command: sct_register_multimodal. This command is designed to register any two images together, so it can be seen as the generalized counterpart to sct_register_to_template.
sct_register_multimodal -i $SCT_DIR/data/PAM50/template/PAM50_t2.nii.gz \
-iseg $SCT_DIR/data/PAM50/template/PAM50_cord.nii.gz \
-d mt1.nii.gz \
-dseg mt1_seg.nii.gz \
-m mask_mt1.nii.gz \
-initwarp ../t2/warp_template2anat.nii.gz \
-param step=1,type=seg,algo=centermass:step=2,type=seg,algo=bsplinesyn,slicewise=1,iter=3 \
-owarp warp_template2mt.nii.gz \
-qc ~/qc_singleSubj
- Input arguments:
-i: Source image. Here, we select the T2 version of the PAM50 template, because the T2 contrast is the closest visual match to our MT1 data.-iseg: Segmented spinal cord for the source image. Here, we use the PAM50 segmented spinal cord volume.-d: Destination image.-dseg: Segmented spinal cord for the destination image.-m: Mask image. This is used on the destination image to improve the accuracy over the region of interest.-initwarp: Warping field used to initialize the source image. Here, we supply thewarp_template2anat.nii.gzfile that was generated in Registering labeled anatomical images to the PAM50 template. Because we start with theTemplate->T2transformation already applied, the resulting warping field will representTemplate->T2->MT1. By comparison, if we registeredTemplate->MT1directly, the warping field could differ from the previous T2 registration. So, specifying-initwarpensures that your registration is more consistent across contrasts.-param: Here, we will tweak the default registration parameters to specify a different nonrigid deformation. The important change isalgo=centermass. Because the template object is already “preregistered” from the previous tutorial (using-initwarp), the benefits of the defaultalgo=centermassrothave already been realized. So, we specify a different algorithm in step 1 to exclude the unnecessary rotation.-owarp: The name of the output warping field. This is optional, and is only specified here to make the output filename a little clearer. By default, the filename would be automatically generated from the filenames-iand-d, which in this case would be the (less clear)warp_PAM50_t22mt1.nii.gz.-qc: Directory for Quality Control reporting. QC reports allow us to evaluate the results slice-by-slice.
- Output files/folders:
PAM50_t2_reg.nii.gz: The PAM50 template image, registered to the space of the MT1 image.warp_template2mt.nii.gz: The warping field to transform the PAM50 template to the MT1 space.
Once the command has finished, at the bottom of your terminal there will be instructions for inspecting the results using Quality Control (QC) reports. Optionally, If you have FSLeyes installed, a fsleyes command will printed as well.