Adding landmark labels for template matching¶
Typically, registering an image to the PAM50 template involves using sct_label_vertebrae to automatically label vertebrae, then selecting 2 intervertebral disc labels to use for matching with the PAM50 template. However, the lumbar region has more variability in cord/disc positioning compared to the cervical and thoracic regions. So, using discs as registration landmarks presents a dilemma: How do we handle variability in the position of the cauda equina relative to the L1-L2 disc?
Notably, in the PAM50 template, the conus medullaris (i.e. the terminal end of the spinal cord) is aligned with the L1-L2 disc. However, for your subjects, the spinal cord may end above or below this point. So, if registration were based on disc landmarks alone, then the tapered region of the spinal cord may end up misaligned with the template.
To correct for this, the PAM50 template provides a cauda equina label (specifically, the tip of the conus medullaris) as a registration landmark. By creating a similar label in your subject data, you can align the terminal end of the spinal cord between your subject and the PAM50 template. However, this comes with the necessary tradeoff of potentially misaligning the discs.
Deciding which landmarks to use (discs, conus medullaris)¶
Registration can be performed with either one or two labels. So, given the tradeoffs between the aligning the discs and/or the conus medullaris, the question becomes: Should registration be done with 1 disc label, 1 conus medullaris label, or a combination of the two?
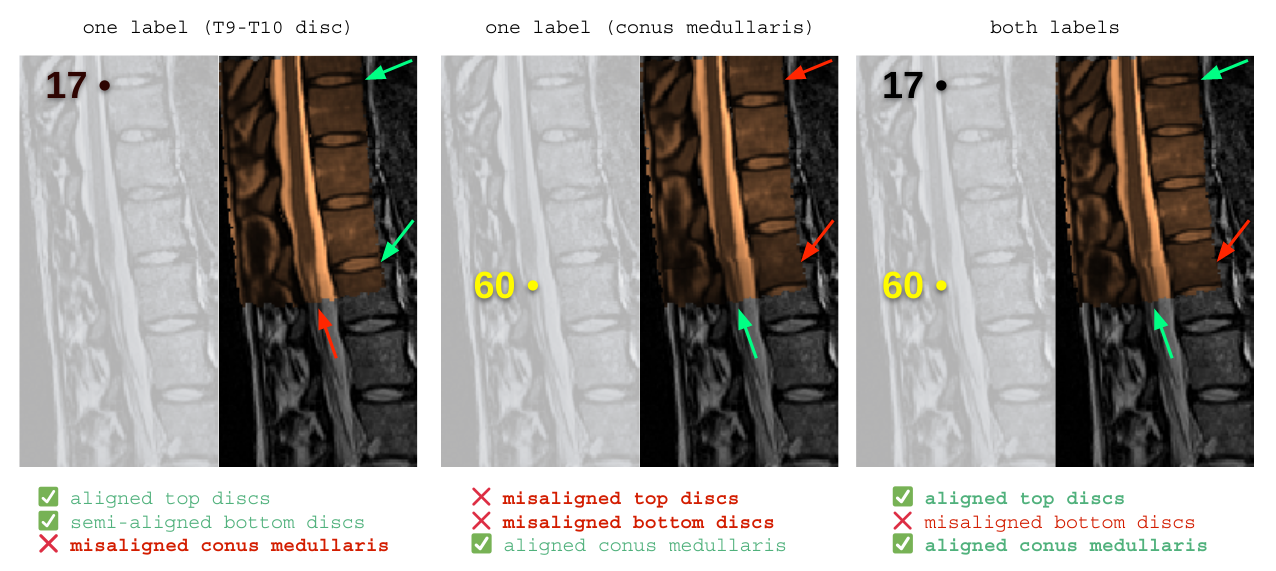
Above is a comparison of sample results, demonstrating that registration will vary significantly depending on which landmarks are chosen. The “ideal result” will depend on the goals of your research. However, in most cases, you will want to stick with one approach and use it consistently throughout your research.
Creating an image with 1 disc label and 1 conus medullaris label¶
As a demonstration, this section will show you how to manually label one disc (T9-T10) as well as the conus medullaris.
There are multiple recommended ways of manually creating labels:
Using
sct_label_utils -create-viewer, which will provide you with a GUI window to select point-wise labels.Determining the coordinates by inspecting the image using a separate 3D image viewer, then adding them via the command line using
sct_label_utils -createUsing the “Edit Mode -> Create Mask” options of a 3D image viewer such as FSLeyes.
Option 1 is the easiest to use when voxel-perfect accuracy is not 100% necessary. However, if you need to precisely locate a specific voxel, we recommend options 2 or 3.
Adding the labels via Option 2: Image Viewer + sct_label_utils -create¶
In this case, voxel accuracy is important, because the conus medullaris label must overlap with the spinal cord segmentation. (This is because of the “straightening step” during registration – the straightening transform is limited to the voxels covered by the spinal cord segmentation, so the registration labels must also exist in that region in order for the labels to be straightened.)
Because of the necessary precision, we recommend that you use Option 2 by following these steps:
Open the anatomical image in an image viewer such as FSLeyes, then overlay the spinal cord segmentation on top.
Move your cursor to the tapered end of the spinal cord.
Adjust the coordinate in all 3 axes to ensure the the cursor is within 0-1 voxel of the edges of the segmentation.
Take note of the coordinate, and provide it to the command below.
For this image, the coordinate [27,79,80] seems to be appropriate location:
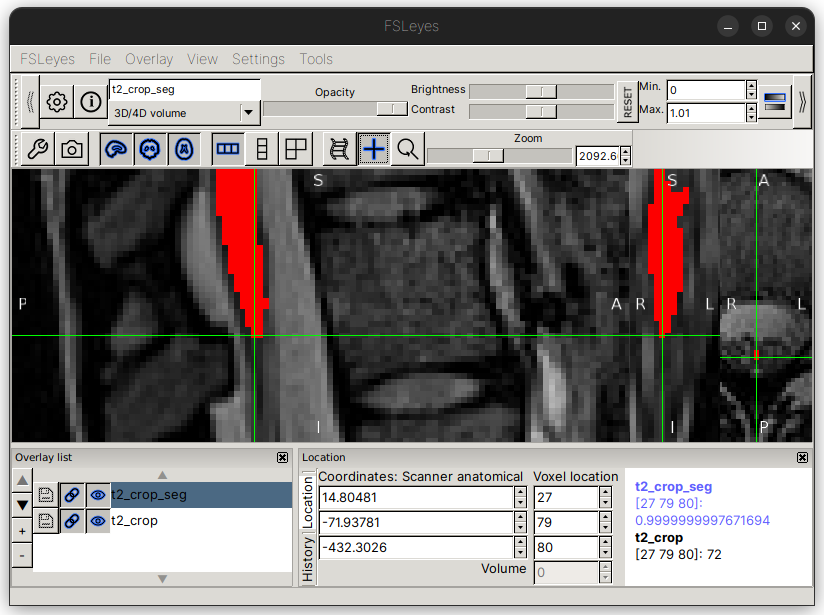
Repeat the same process for the posterior tip of the T9-T10 disc. Here, we will use the coordinate [22,77,187]:
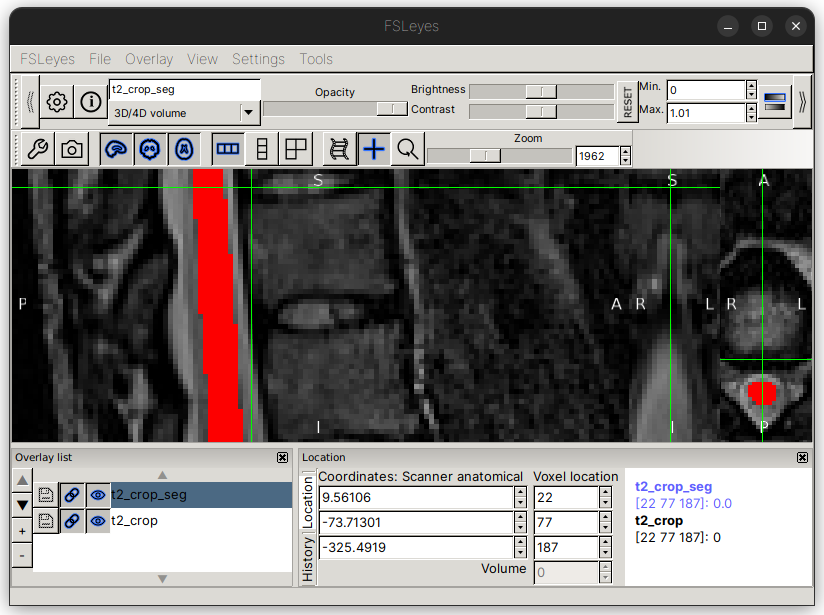
The command below will add labels for the T9-T10 disc (numerical ID 17) and the conus medullaris (numerical ID 60).
sct_label_utils -i t2_lumbar.nii.gz -create 22,77,187,17:27,79,80,60 -o t2_lumbar_labels.nii.gz
- Input arguments:
-i: The input anatomical image.-create: This argument will create a label with value 17 (T9-T10 disc) at coordinate[22,77,187], and a label with value 60 (conus medullaris) at the coordinate[27,79,80].-o: The name of the output file.
- Output files/folders:
t2_lumbar_labels.nii.gz: An image containing two single-voxel labels.
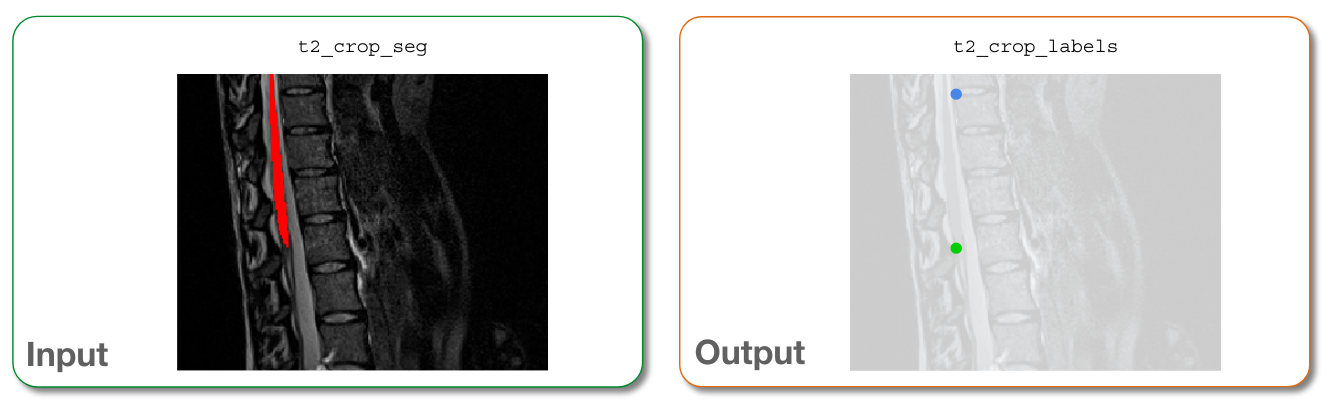

Input/output images for sct_label_utils -create¶