Alternative #2: Manual labeling all labels¶
In special circumstances, both the automated and semi-automated approaches may not be applicable. For example, the C2-C3 disc may not be present in your image, or you may be dealing with heavily artifacted data. In those cases, you may want to skip the automated labeling approaches, and instead manually label every vertebrae.
As before, you can use the -create-viewer option, this time providing multiple values. For example, if you wish to label the C2-C3, C3-C4, and C4-C5 discs, you would run the following command:
sct_label_utils -i t2.nii.gz -create-viewer 3,4,5 -o labels_disc.nii.gz \
-msg "Place labels at the posterior tip of each inter-vertebral disc. E.g. Label 3: C2/C3, Label 4: C3/C4, etc."
- Input arguments:
-i: The input anatomical image.-create-viewer: This argument will open up an interactive GUI coordinate picker that will prompt you to label the discs corresponding to the IDs3,4,5(C2-C3, C3-C4, and C4-C5 discs respectively).-msg: The text that will appear at the top of the window.-o: The name of the output file.
- Output files/folders:
labels_disc.nii.gz: An imagine containing the single-voxel label as selected in the GUI coordinate picker.
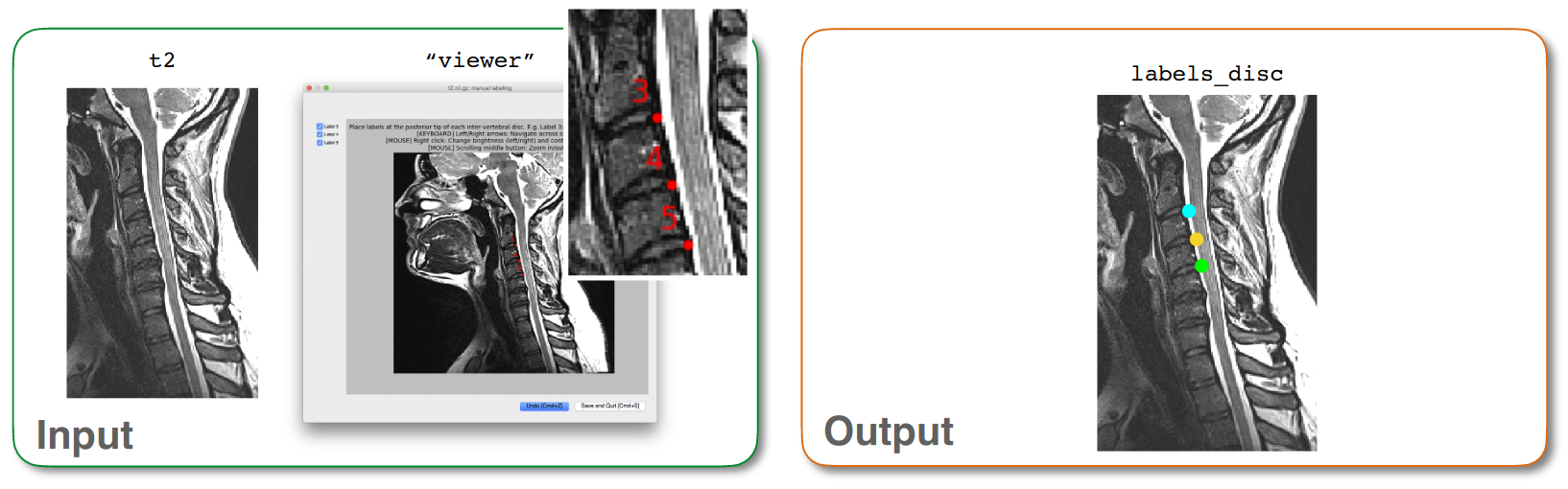
Input/output images for sct_label_utils -create-viewer 3,4,5¶
Note
We recommend that you try labeling inteverbral discs as opposed to vertebral bodies, as it is often easier to accurately select the posterior tip of the disc with a mouse pointer.